 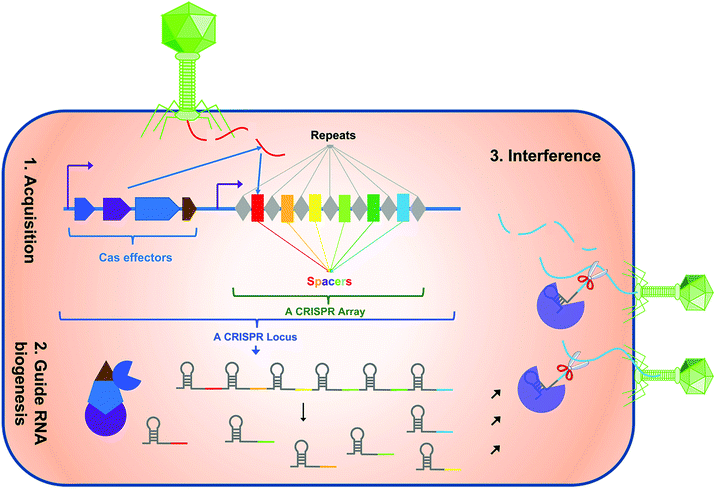
The species with the largest number of genomes with CRISPR-Cas systems were K. aureus despite having started from a larger number of genomes ( Fig. According to the number of genes, the smallest pangenome was found for S. 
The genomes of the different ESKAPE species were initially obtained, and they were both structurally and functionally annotated with a special emphasis on the protein-coding genes and the elements that are part of the CRISPR-Cas systems. RESULTS A large proportion of CRISPR dark matter spacers can be annotated by pangenome analysis In addition, our results demonstrate that the study of thousands of genomes of the same species allows us to reduce the CRISPR dark matter and to trace the origin of most of the spacers found in them. This has motivated us to investigate whether this relationship could be mediated by phages that could take advantage of the existence of these membrane proteins as receptors or adhesion sites to infect bacteria. Then, these genes have been functionally analyzed, and we found that they are enriched in genes encoding membrane proteins. We have created a large pangenome for each species and compared strains with or without CRISPR-Cas systems to discover genes associated with the first group. ESKAPE bacteria have CRISPR-Cas systems of the most common types, from I to IV, but only in a minimal number of strains, with frequencies ranging from less than 1 to 60% of genomes, depending on the species ( 16). In this work, we analyzed thousands of genomes of the group of bacteria known as ESKAPE, whose acronym refers to two Gram-positive bacteria ( Enterococcus faecium and Staphylococcus aureus) and four Gram-negative bacteria ( Klebsiella pneumoniae, A. There are different types of CRISPR-Cas systems based on genes that are part of the different steps of this immune system (adaptation or spacer integration, expression, and interference) and are generically called cas (CRISPR-associated) genes. They provide acquired immune resistance against phages and other foreign nucleic acid molecules such as plasmids, thus restricting gene transfer ( 4). CRISPR-Cas is an adaptive immunity system found in most archaea and in less than half of the bacteria sequenced ( 3). Restriction-modification systems are by far the most abundant, being present in 83% of prokaryotic genomes, followed by CRISPR-Cas with about 40% ( 2). Bacteria defend themselves against infection by these phages by means of different molecular strategies. They can be used to control bacterial growth and are even beginning to be used to treat infections in humans ( 1). To our knowledge this is the first example of gene therapy for an autosomal dominant gain-of-function disease using a disease allele inactivation strategy in place of the less efficient disease allele repair approach.Bacteriophages, also known as phages, are viruses that predate bacterial cells, representing environmental burdens for their growth and spread. The WHIM allele-inactivated HSCs retained long-term pluripotency and selective hematopoietic reconstitution advantages. Then, through in vivo natural selection, WHIM allele-inactivated cells were enriched over wildtype allele-inactivated cells. First, one copy of Cxcr4 in HSPCs was inactivated in vitro by CRISPR/Cas9 editing with a single guide RNA (sgRNA) that does not discriminate between WHIM and wildtype Cxcr4 alleles. We have now developed a two-step preclinical protocol of autologous hematopoietic stem and progenitor cell (HSPC) transplantation towards this goal. We previously reported that Cxcr4-haploinsufficient ( Cxcr4 +/o) hematopoietic stem cells (HSCs) have a strong selective advantage for durable hematopoietic reconstitution over wildtype ( Cxcr4 +/+) and WHIM ( Cxcr4 +/w) HSCs, and that a WHIM patient was spontaneously cured by chromothriptic deletion of the disease allele in an HSC, suggesting that WHIM allele inactivation through gene editing may be a safe genetic cure strategy for the disease. WHIM syndrome is an autosomal dominant immunodeficiency disorder caused by gain-of-function mutations in chemokine receptor CXCR4 that promote severe panleukopenia due to bone marrow (BM) retention of mature leukocytes. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
